MRI源文件
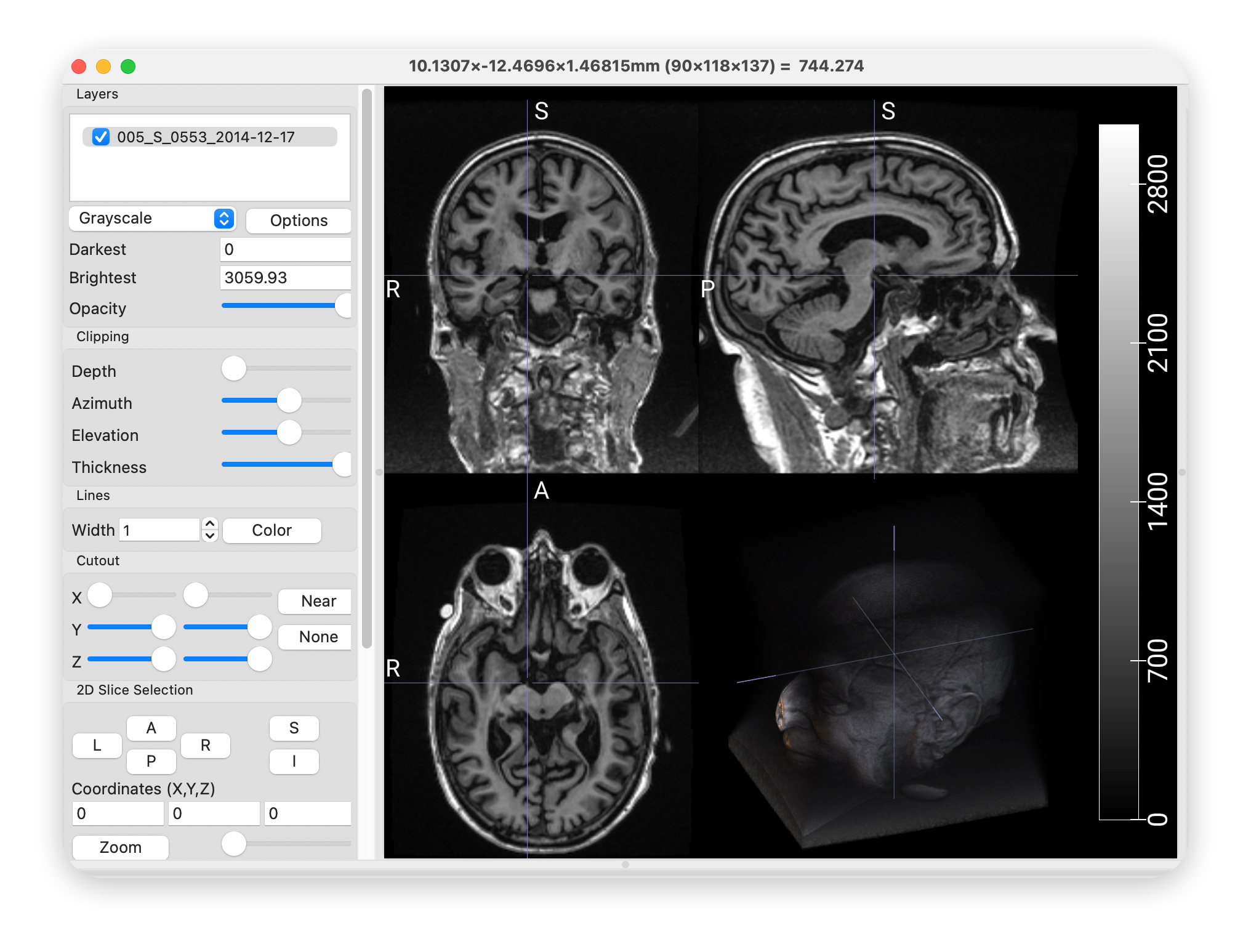
step 1 颅骨去除
1.依赖安装-HD-BET
bash
pip install hd-bet2.单一文件
bash
hd-bet -i INPUT_FILENAME -o OUTPUT_FILENAME3.多个文件
bash
hd-bet -i INPUT_FOLDER -o OUTPUT_FOLDER4.输出图像
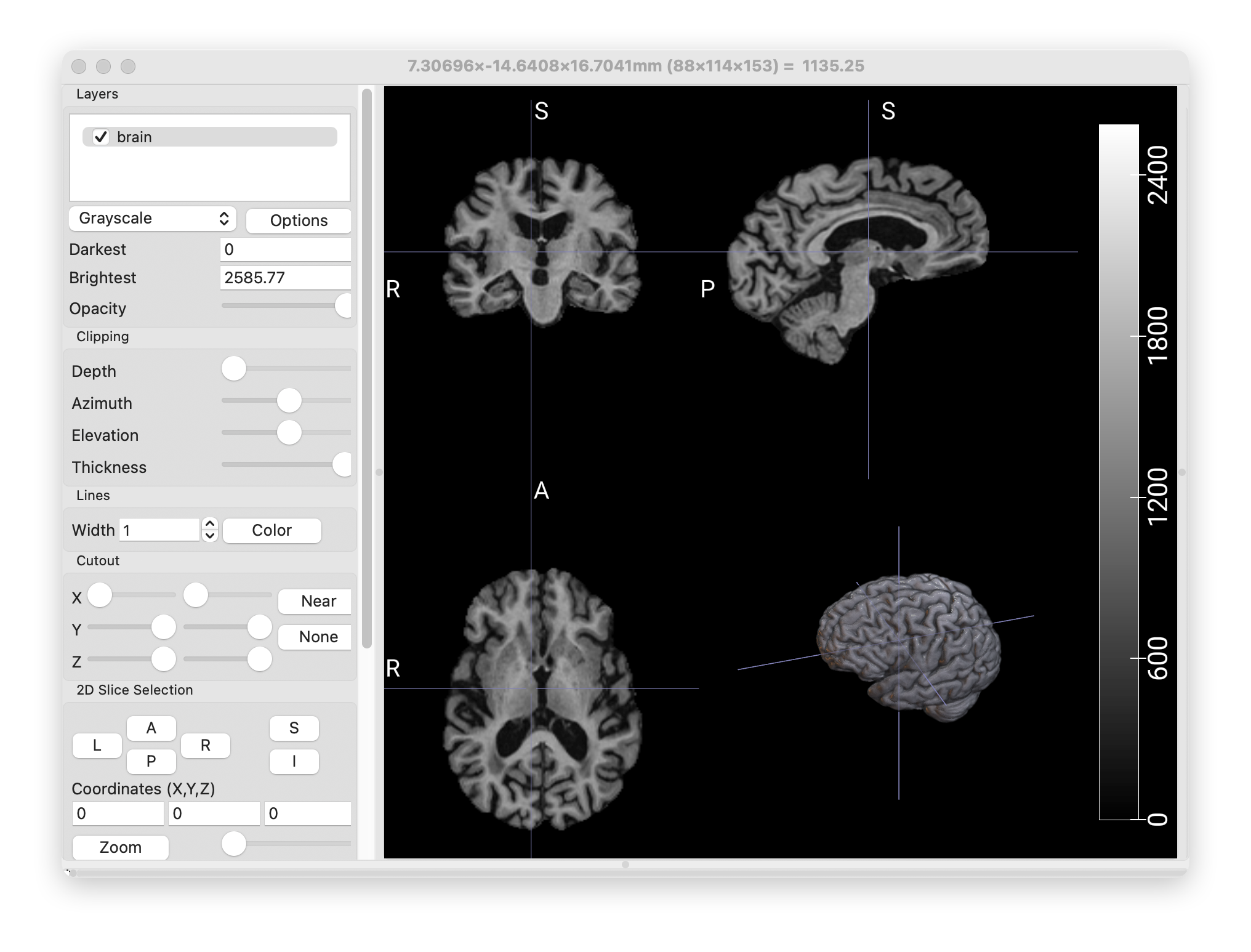
step 2 离线增强
1.依赖安装-ANTsPy
bash
pip install antspyx2.图像读取
python
img = ants.image_read(input_file)3.N4偏置场校正
同一种组织,在图像不同位置的灰度可能不一致,解决亮度不均匀问题。
python
mask = ants.get_mask(img)
img = ants.n4_bias_field_correction(img, mask=mask)4.图像方向统一RAS
不同设备、不同医院导出的 MRI 数据,其坐标方向可能不同。 LPS、LAS、RAS。
python
img = ants.reorient_image2(img, orientation="RAS")5.重采样
不同 MRI 数据的体素大小可能不同,可根据具体任务显存占用大小进行调节,越小占用越大
python
img = ants.resample_image(img, (1.0, 1.0, 1.0), use_voxels=False, interp_type=1)6.配准到标准模板MNI
不同受试者的脑结构位置仍然不一致,需要将所有图像配准到同一个标准模板空间。
text
reg = ants.registration(fixed=template, moving=img, type_of_transform="Affine")
img_reg = reg["warpedmovout"]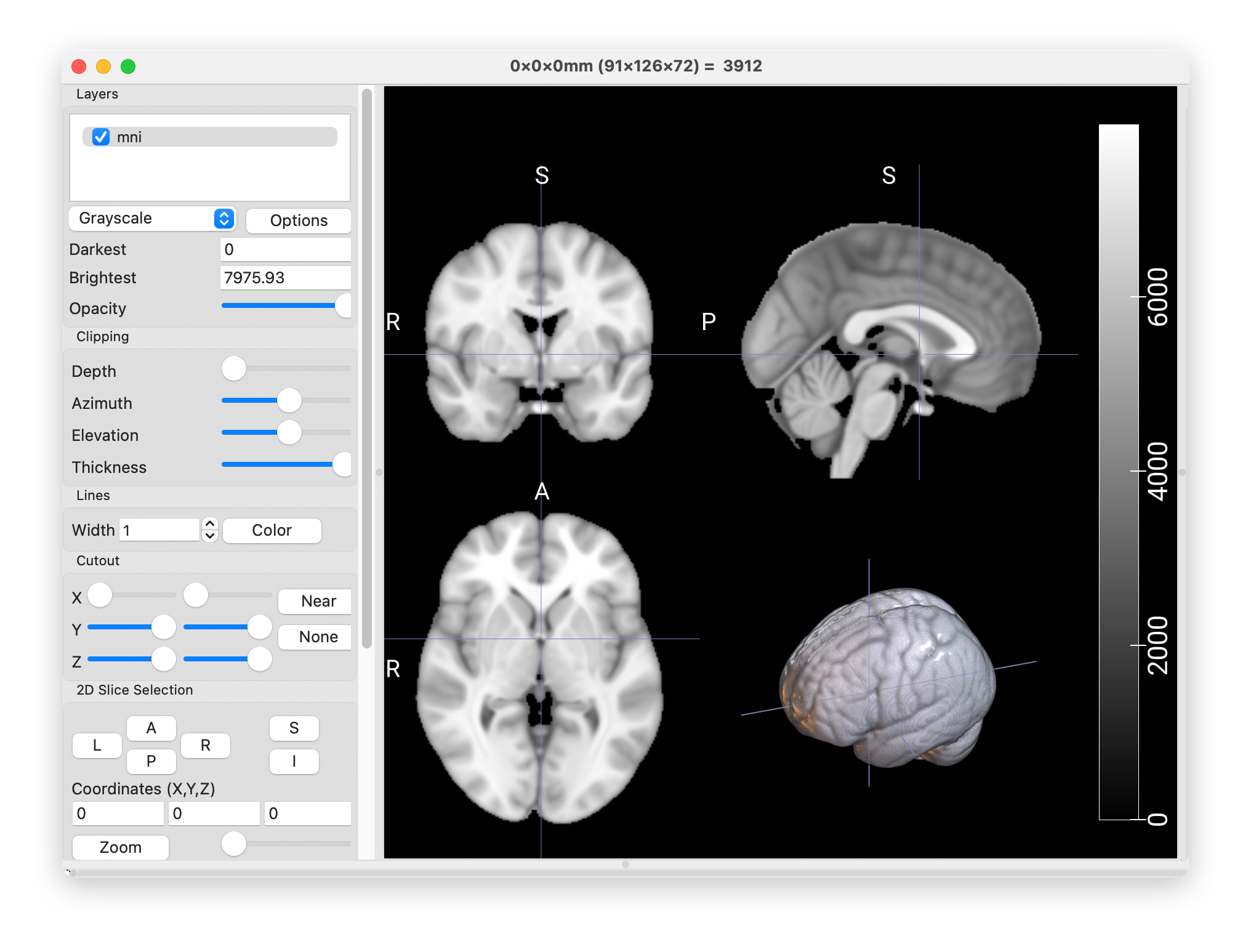
7.结果保存和变换矩阵
保存变换文件有几个重要用途:
- 后续把分割标签映射到模板空间
- 把模板空间结果反变换回原始空间
- 复现处理流程
- 检查每个样本的配准参数
python
ants.image_write(img_reg, output_file)
for i, t in enumerate(reg["fwdtransforms"]):
ext = os.path.splitext(t)[1]
save_path = os.path.join(transform_dir, f"{base_name}_fwd_{i}{ext}")
shutil.copy(t, save_path)
for i, t in enumerate(reg["invtransforms"]):
ext = os.path.splitext(t)[1]
save_path = os.path.join(transform_dir, f"{base_name}_inv_{i}{ext}")
shutil.copy(t, save_path)8.输出图像
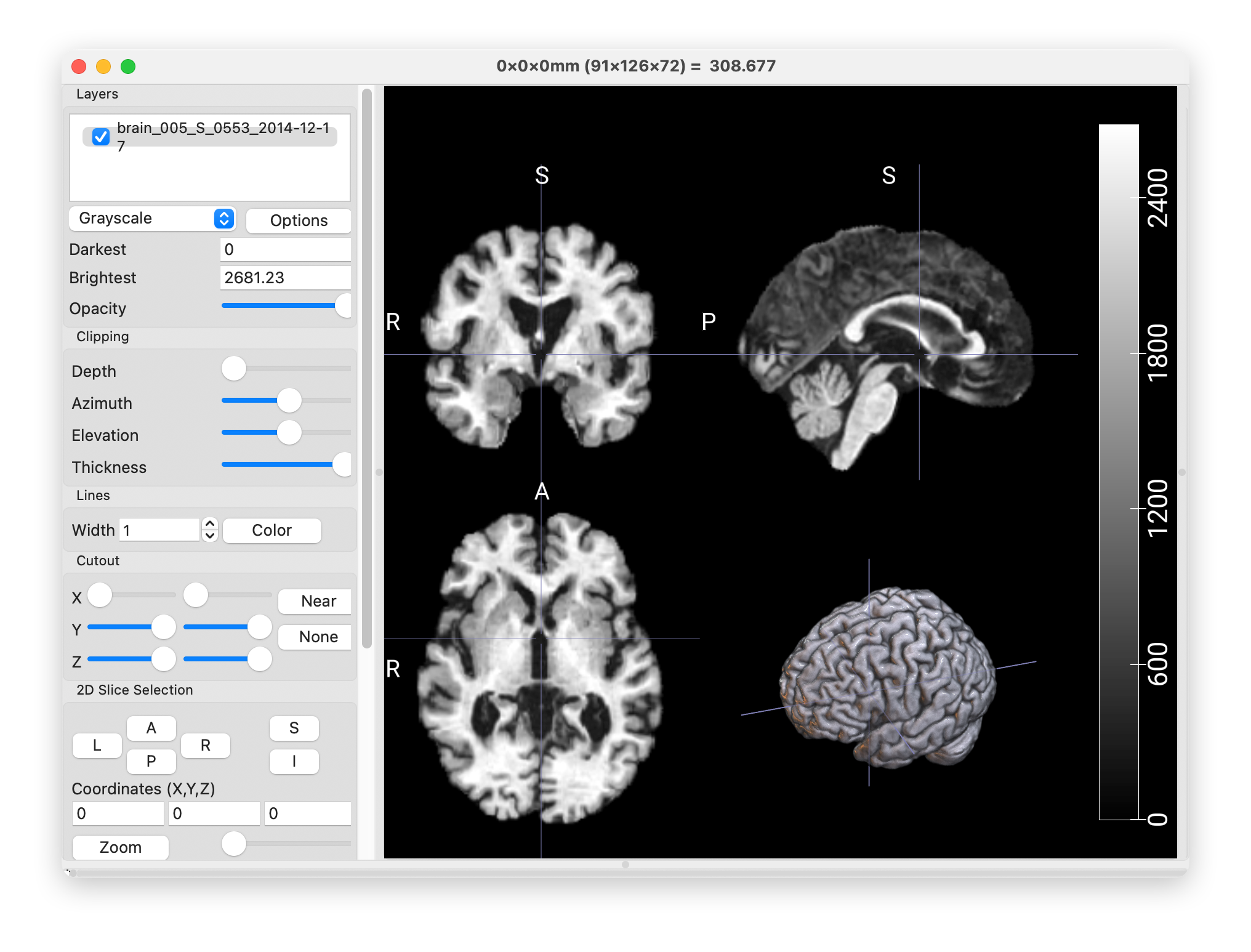
9.完整代码
python
import ants
import os
import shutil
def preprocess(input_file, output_file, template, transform_dir=None):
img = ants.image_read(input_file)
# 1. N4
mask = ants.get_mask(img)
img = ants.n4_bias_field_correction(img, mask=mask)
# 2. RAS
img = ants.reorient_image2(img, orientation="RAS")
# 3. resample spacing
img = ants.resample_image(img, (1.0, 1.0, 1.0), use_voxels=False, interp_type=1)
# 4. registration
reg = ants.registration(fixed=template, moving=img, type_of_transform="Affine")
img_reg = reg["warpedmovout"]
# 5. save registered image
ants.image_write(img_reg, output_file)
# 6. save transforms
if transform_dir is not None:
os.makedirs(transform_dir, exist_ok=True)
base_name = os.path.basename(input_file)
if base_name.endswith(".nii.gz"):
base_name = base_name[:-7]
elif base_name.endswith(".nii"):
base_name = base_name[:-4]
for i, t in enumerate(reg["fwdtransforms"]):
ext = os.path.splitext(t)[1]
save_path = os.path.join(transform_dir, f"{base_name}_fwd_{i}{ext}")
shutil.copy(t, save_path)
for i, t in enumerate(reg["invtransforms"]):
ext = os.path.splitext(t)[1]
save_path = os.path.join(transform_dir, f"{base_name}_inv_{i}{ext}")
shutil.copy(t, save_path)
if __name__ == "__main__":
input_dir = "/datasets/GPH-TCM/nii_data/MRI-T1_only_brain/"
output_dir = "/datasets/GPH-TCM/nii_data/MRI-T1_only_brain_preprocessed/"
transform_dir = "/datasets/GPH-TCM/nii_data/MRI-T1_only_brain_transforms/"
template_path = " mni.nii.gz"
os.makedirs(output_dir, exist_ok=True)
os.makedirs(transform_dir, exist_ok=True)
template = ants.image_read(template_path)
for filename in os.listdir(input_dir):
if filename.endswith(".nii.gz") or filename.endswith(".nii"):
input_file = os.path.join(input_dir, filename)
output_file = os.path.join(output_dir, filename)
try:
preprocess(input_file, output_file, template, transform_dir)
print(f"[OK] {filename}")
except Exception as e:
print(f"[ERROR] {filename}: {e}")
Step 3 在线增强
python
from monai.transforms import (
Compose,
EnsureChannelFirstd,
EnsureTyped,
Lambdad,
LoadImaged,
NormalizeIntensityd,
RandScaleIntensityd,
RandShiftIntensityd,
)
# train
train_transform = Compose(
[
LoadImaged(keys="image"),
EnsureChannelFirstd(keys="image"),
NormalizeIntensityd(keys="image", nonzero=True),
RandScaleIntensityd(keys="image", factors=0.1, prob=0.3),
RandShiftIntensityd(keys="image", offsets=0.1, prob=0.3),
EnsureTyped(keys=["image"]),
Lambdad(keys="labels", func=lambda x: torch.tensor(x, dtype=torch.long)),
]
)
# val or test
val_transform = Compose(
[
LoadImaged(keys="image"),
EnsureChannelFirstd(keys="image"),
NormalizeIntensityd(keys="image", nonzero=True),
EnsureTyped(keys=["image"]),
Lambdad(keys="labels", func=lambda x: torch.tensor(x, dtype=torch.long)),
]
)