使用说明
该脚本完成以下工作:
- 加载 3D NIfTI 医学影像(如 MRI)
- 使用 MONAI 进行预处理(需调整为训练所用的预处理)
- 使用训练好的模型进行推理
- 通过 SmoothGrad 生成梯度注意力图
- 对 heatmap 进行后处理(平滑 + mask + 归一化)
- 输出 .nii.gz 热力图文件及相关信息
输出示例
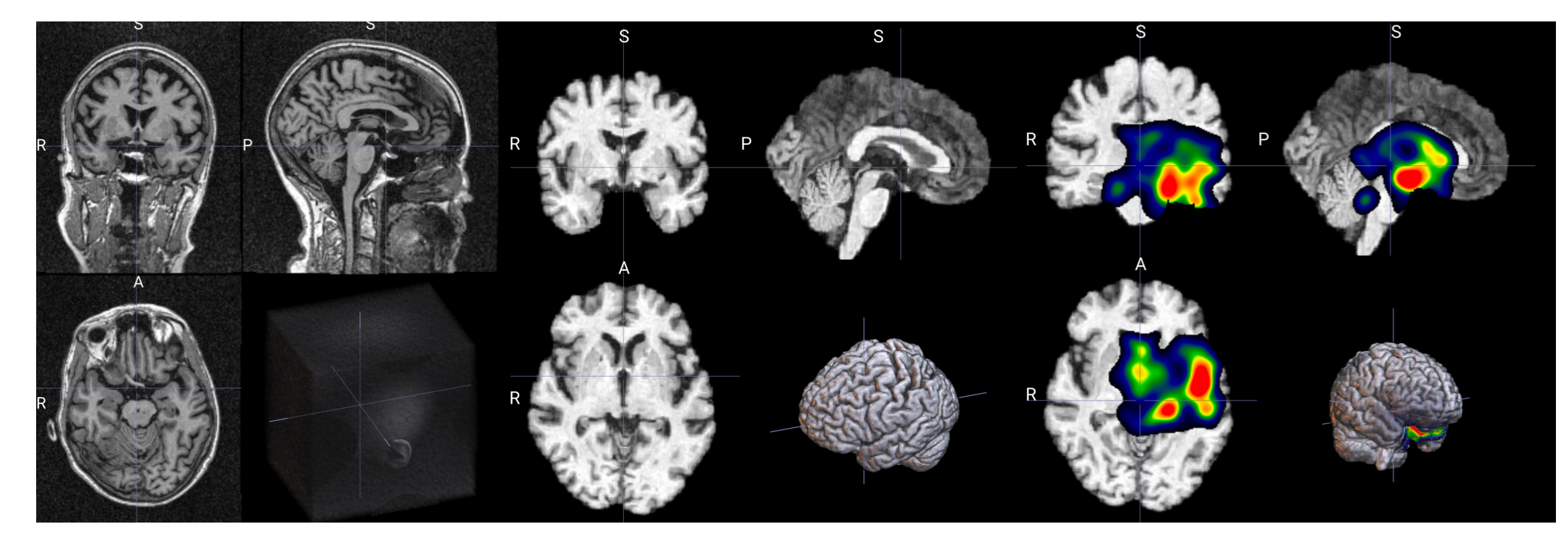
所需依赖
bash
pip install nibabel numpy scipy torch monai参数说明
模型相关
--model_name
模型名称用于导入模型 参考如下
python
def select_model(args):
model_name = args.model_name
num_classes = args.num_classes
if model_name == "SegTom":
return SegTom(in_channels=1, num_classes=num_classes)
# elif model_name == "nnMamba":
# return nnMambaEncoder(number_classes=num_classes)
elif model_name == "MedMamba":
return MobileNetV2(num_classes=num_classes, in_channels=1, alpha=0.75)
elif model_name == "EfficientNet":
return EfficientNetBN(
spatial_dims=3,
in_channels=1,
out_channels=num_classes,
alpha=0.75,
)
elif model_name == "DenseNet169":
return DenseNet169(spatial_dims=3, in_channels=1, out_channels=num_classes)
elif model_name == "DenseNet264":
return DenseNet264(spatial_dims=3, in_channels=1, out_channels=num_classes)
elif model_name == "ResViT":
return ResViTClassifier(num_classes=num_classes)
elif model_name == "RDNet":
return RDNet(num_classes=num_classes)
else:
raise ValueError(f"Unknown model name: {model_name}")--num_classes
分类类数,用于加载模型
--model_path
模型权重路径
--nii_name
输入文件名
--origin_nii_file
原始文件夹路径,用于将原文件复制到outputs,为空则不复制
--brain_nii_file
预处理后文件夹路径,即训练数据集
--nii_label
指定解析标签,用于选定需要解释的类,为-1则按推理结果解释
系统相关
--output_dir
输出文件夹目录
为空则默认输出格式为 output_dir/nii_name/.....
--device
使用设备
核心参数
--smoothgrad_samples 实测区别不大
控制 SmoothGrad 采样次数
- 给输入图像加一点随机噪声
- 每次都做一次前向传播和反向传播
- 把多次得到的梯度结果累加平均
bash
--smoothgrad_samples 8 # 快但噪声大
--smoothgrad_samples 24 # 推荐
--smoothgrad_samples 48 # 最稳定但慢--smoothgrad_noise_std
控制每次给输入图像添加多少噪声。
噪音小:
- 更贴近原始梯度
- 细节更多
- 但噪声抑制不明显
- 热力图可能会碎
噪音大:
- 更平滑
- 更偏“大范围响应”
- 细节会被抹掉
- 甚至可能出现关注区域漂移
--smoothgrad_noise_std 0.8
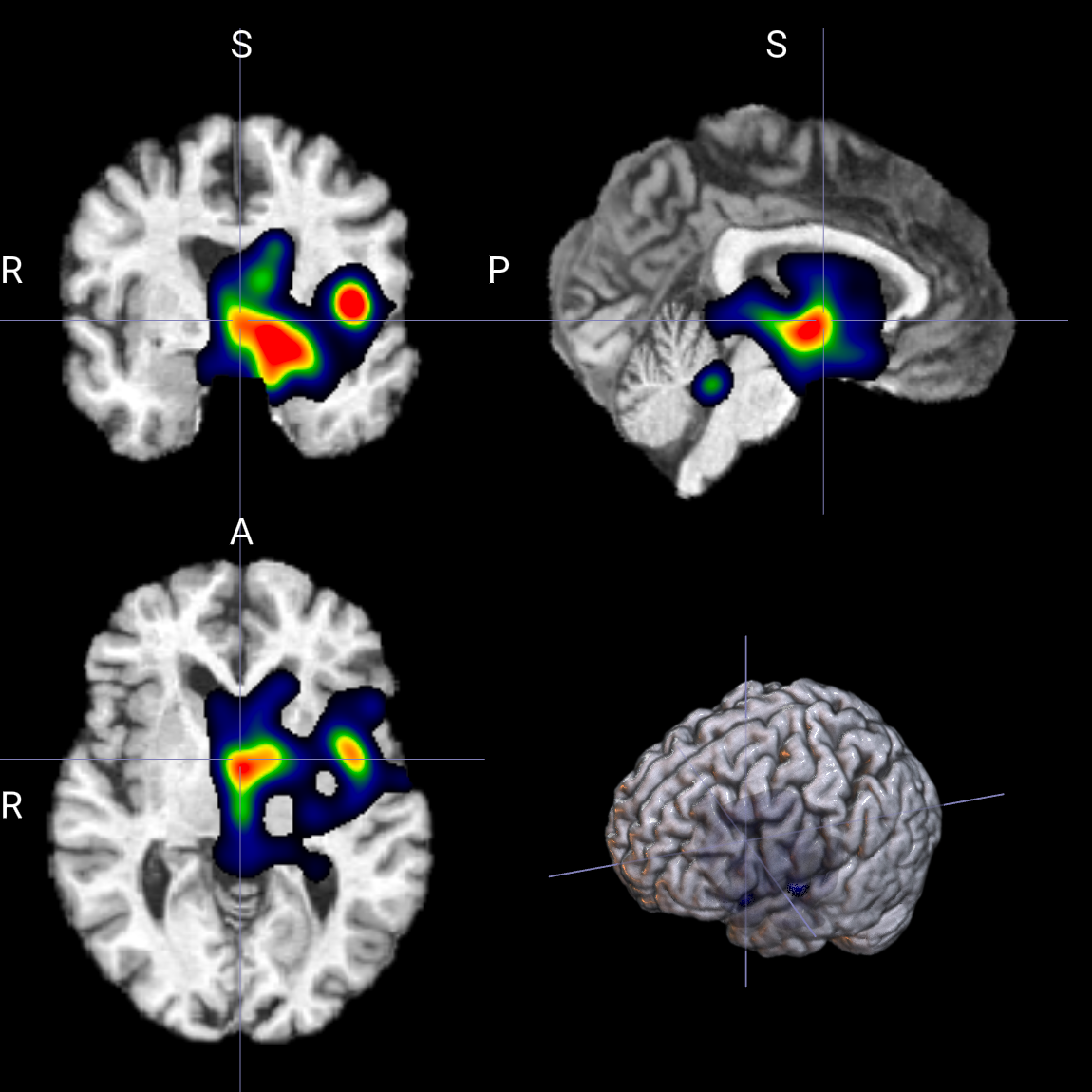
--smoothgrad_noise_std 10
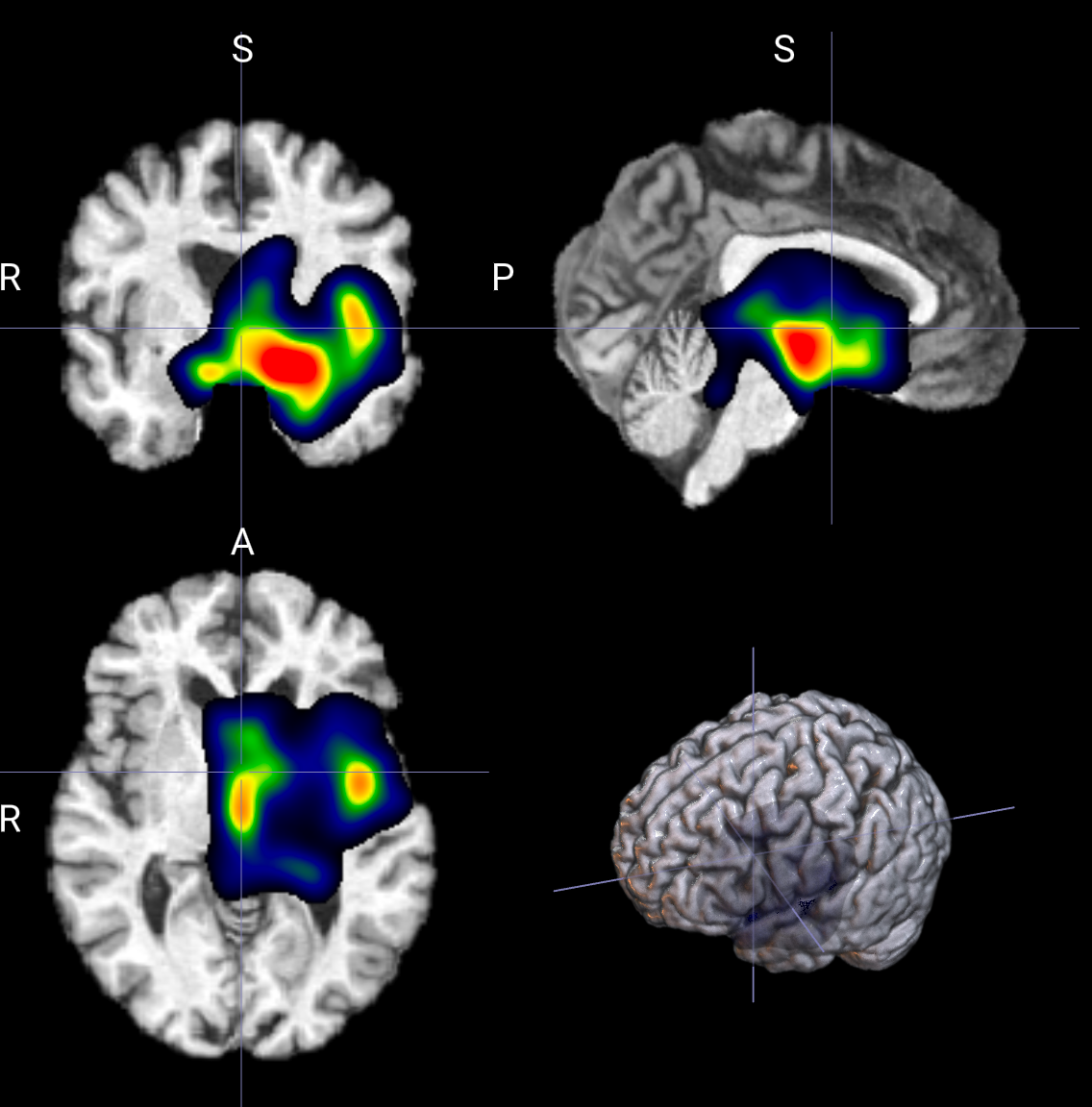
--nii_label
控制想解释模型对哪个类别的响应。
图像中哪些区域,对某个类别的分数最有贡献。
模式 1:解释模型当前预测
text
--nii_label -1这时逻辑是:
- 模型先预测一个类别
- 再解释“为什么模型会认为它属于这个类”
这是最常见的解释方式。
适合问题:
- 模型为什么判成 AD?
- 模型为什么把它分到第 2 类?
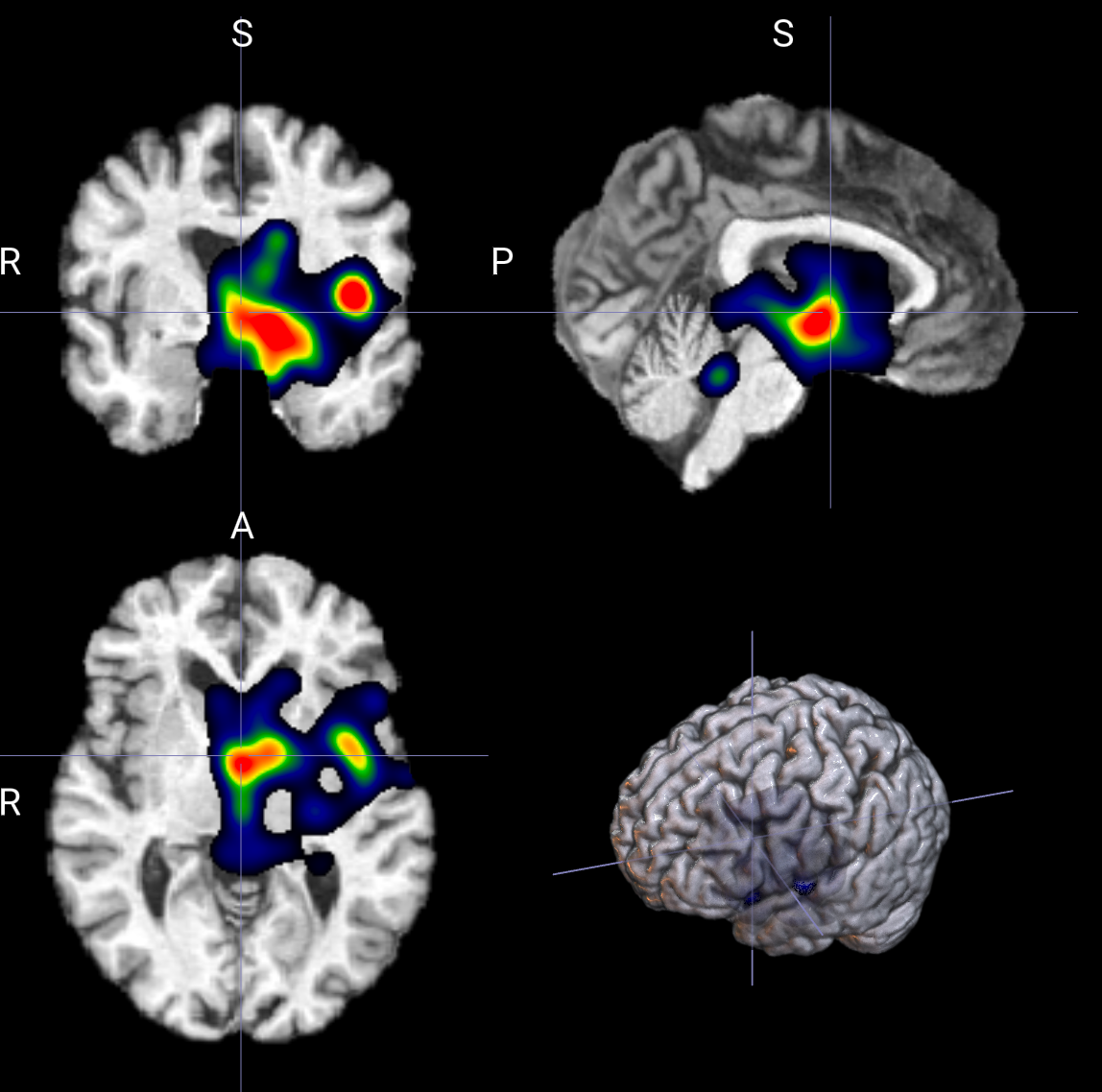
模式 2:解释指定类别
text
--nii_label 2这时逻辑是:
- 不管模型预测了什么
- 我都要看“对类别 1 的支持证据在哪里”
适合问题:
- 即使模型没判成 AD,我也想看 AD 相关区域有没有激活
- 做类间对比解释
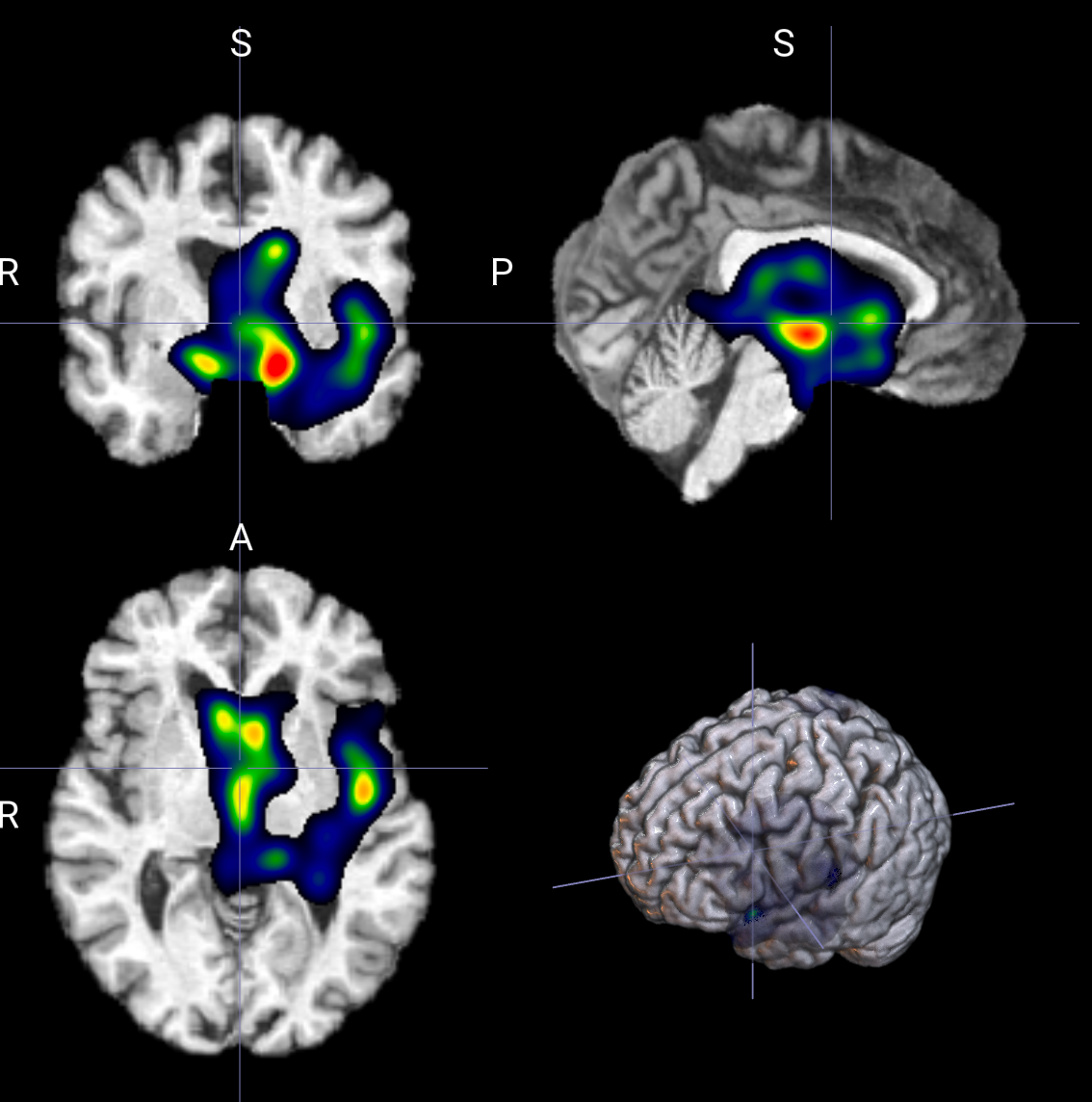
--display_downsample
控制 显示重建前的降采样因子,主要用于让 heatmap 更平滑、更像连续区域,而不是零碎散点
- 先把热图缩小
- 在低分辨率空间做平滑
- 再放大回来
bash
--display_downsample 1 # 表示不做降采样重建。
--display_downsample 2 # 通常比较自然 能去掉一部分毛刺 仍保留结构边界
--display_downsample 4 # 更强的结构化平滑 区域更块状 细节会明显减少--display_downsample 1
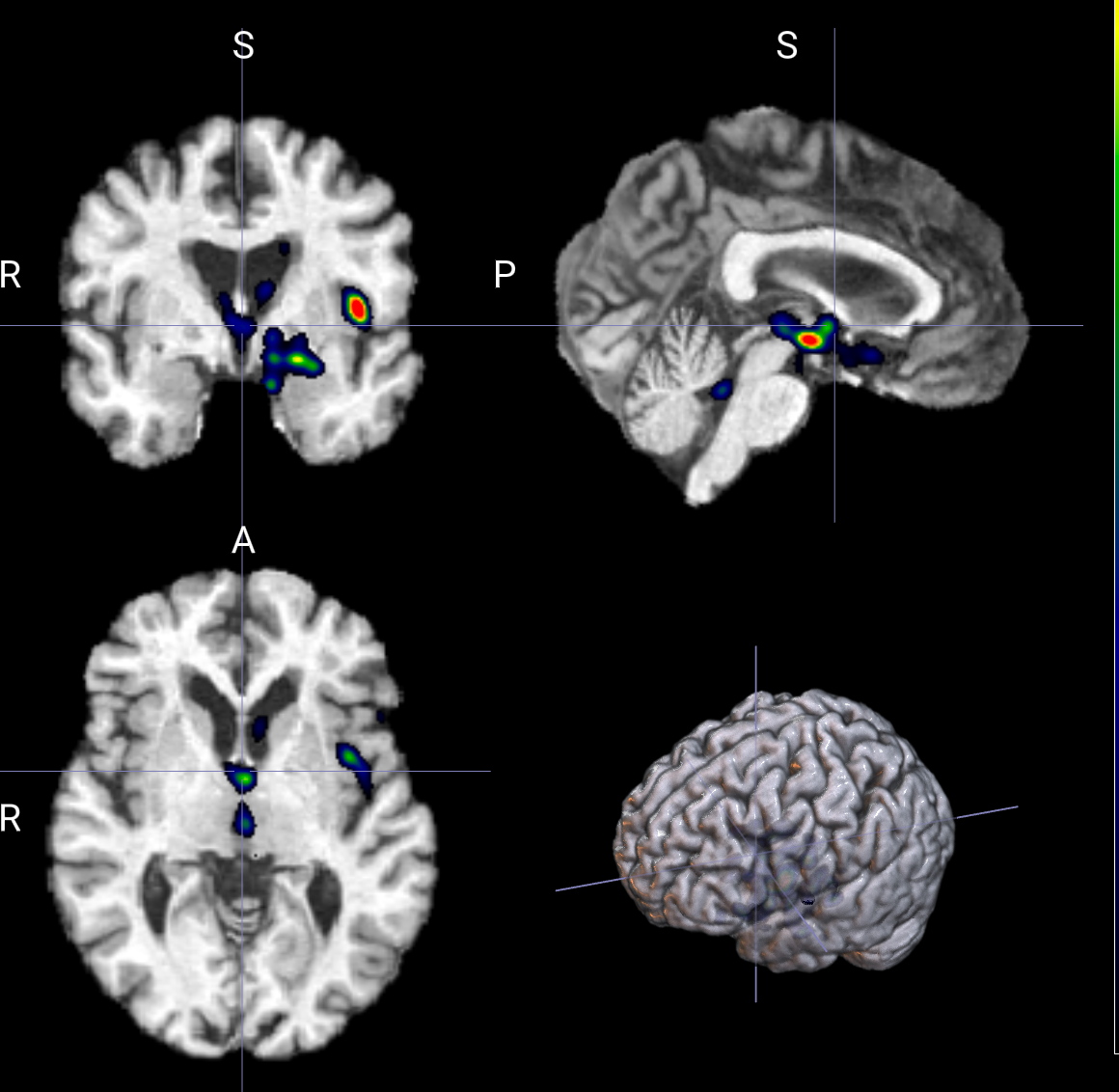
--display_downsample 4
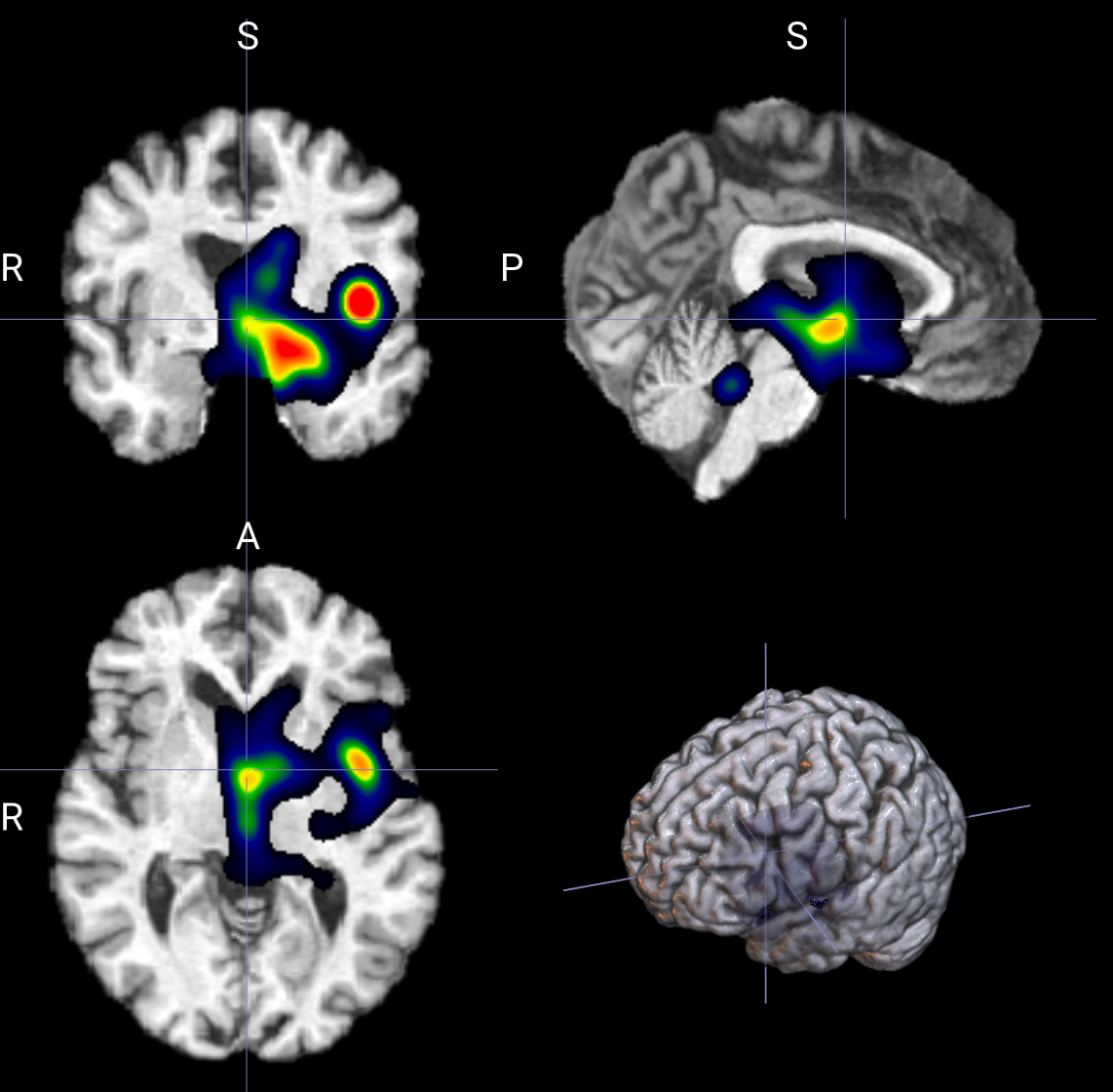
--display_blur_sigma 实测区别不大
控制最后输出热图时的 高斯模糊强度。
bash
--display_blur_sigma 0.8 # 更锐利 保留更多局部形状 但可能更毛躁
--display_blur_sigma 1.6 # 平滑和细节平衡较好 视觉上更自然
--display_blur_sigma 3.0 # 很平滑 小热点会被融掉 更适合做大区域展示--display_supersample 实测区别不大
控制 超采样插值平滑
- 先把热图放大
- 在高分辨率上平滑
- 再缩回原尺寸
bash
--display_supersample 1 # 不做超采样
--display_supersample 2 # 通常
--display_supersample 3 # 更细腻,但计算更多--display_clip_min 和 --display_clip_max
意思是:
- 小于
clip_min的值,统统压成clip_min - 大于
clip_max的值,统统压成clip_max - 然后重新映射到
[0,1]
多弱的响应会被当成“背景”压掉。
多强的响应会被压平,防止极亮点支配整张图。
使用demo
默认demo
bash
python hotmap.py \
--device cuda:0 \
--nii_name 037_S_0454_2015-07-15.nii.gz \
--origin_nii_file /datasets/GPH-TCM/nii_data/MRI-T1_origin/ \
--brain_nii_file /datasets/GPH-TCM/nii_data/MRI-T1/ \
--nii_label -1 \
--smoothgrad_samples 24 \
--smoothgrad_noise_std 0.14 \
--display_downsample 4 \
--display_blur_sigma 1.6 \
--display_supersample 2 \
--display_clip_min 0.2 \
--display_clip_max 0.8方案一:快速调试型
适合先跑通流程,看大概效果。
bash
--smoothgrad_samples 8 \
--smoothgrad_noise_std 0.10 \
--display_downsample 4 \
--display_blur_sigma 1.6 \
--display_supersample 1 \
--display_clip_min 0.10 \
--display_clip_max 0.90特点:
- 快
- 图比较干净
- 细节不是重点
方案二:平衡通用型
适合大多数实验。
bash
--smoothgrad_samples 24 \
--smoothgrad_noise_std 0.14 \
--display_downsample 2 \
--display_blur_sigma 1.6 \
--display_supersample 2 \
--display_clip_min 0.10 \
--display_clip_max 0.90特点:
- 稳定
- 细节和可读性平衡
- 推荐默认
方案三:细节保留型
适合想看更细局部结构。
bash
--smoothgrad_samples 32 \
--smoothgrad_noise_std 0.08 \
--display_downsample 1 \
--display_blur_sigma 0.8 \
--display_supersample 2 \
--display_clip_min 0.05 \
--display_clip_max 0.95特点:
- 更锐利
- 更容易看到局部差异
- 但可能更噪
方案四:论文展示型
适合做最终图。
bash
--smoothgrad_samples 48 \
--smoothgrad_noise_std 0.12 \
--display_downsample 2 \
--display_blur_sigma 1.2 \
--display_supersample 3 \
--display_clip_min 0.08 \
--display_clip_max 0.88特点:
- 稳定
- 边界更自然
- 观感更好
完整代码
python
import argparse
import copy
import os
import shutil
from typing import Tuple
import nibabel as nib
import numpy as np
import scipy.ndimage as ndi
import torch
from monai.transforms import (
Compose,
EnsureChannelFirstd,
EnsureTyped,
LoadImaged,
NormalizeIntensityd,
)
from src.utils import select_model
val_transform = Compose(
[
LoadImaged(keys="image"),
EnsureChannelFirstd(keys="image"),
NormalizeIntensityd(keys="image", nonzero=True),
EnsureTyped(keys=["image"]),
]
)
def _strip_nii_ext(filename: str) -> str:
if filename.endswith(".nii.gz"):
return filename[:-7]
return os.path.splitext(filename)[0]
def _normalize_01(arr: np.ndarray, eps: float = 1e-8) -> np.ndarray:
arr = np.asarray(arr, dtype=np.float32)
min_v = float(arr.min())
max_v = float(arr.max())
if max_v - min_v < eps:
return np.zeros_like(arr, dtype=np.float32)
return (arr - min_v) / (max_v - min_v + eps)
def _build_brain_mask(brain_arr: np.ndarray) -> np.ndarray:
mask = np.asarray(brain_arr > 0, dtype=bool)
mask = ndi.binary_fill_holes(mask)
# Light morphology to suppress isolated noise while preserving inner structures.
mask = ndi.binary_opening(mask, structure=np.ones((3, 3, 3), dtype=bool))
return mask
def _postprocess_attention(attn01: np.ndarray, brain_mask: np.ndarray) -> np.ndarray:
smooth = ndi.gaussian_filter(attn01, sigma=1.0)
masked = smooth * brain_mask.astype(np.float32)
nonzero_vals = masked[brain_mask]
if nonzero_vals.size == 0:
return np.zeros_like(masked, dtype=np.float32)
low_q = float(np.quantile(nonzero_vals, 0.45))
high_q = float(np.quantile(nonzero_vals, 0.90))
denom = max(high_q - low_q, 1e-8)
soft = np.clip((masked - low_q) / denom, 0.0, 1.0)
soft = np.power(soft, 1.35)
support = soft > 0.10
support &= brain_mask
support = ndi.binary_closing(support, structure=np.ones((3, 3, 3), dtype=bool), iterations=2)
support = ndi.binary_opening(support, structure=np.ones((2, 2, 2), dtype=bool), iterations=1)
support_soft = ndi.gaussian_filter(support.astype(np.float32), sigma=0.8)
refined = soft * np.clip(support_soft, 0.0, 1.0)
refined *= brain_mask.astype(np.float32)
return _normalize_01(refined)
def _merge_for_display(
attn01: np.ndarray,
brain_mask: np.ndarray,
blur_sigma: float,
supersample: int,
) -> np.ndarray:
base = attn01 * brain_mask.astype(np.float32)
ms1 = ndi.gaussian_filter(base, sigma=1.2)
ms2 = ndi.gaussian_filter(base, sigma=2.0)
ms3 = ndi.gaussian_filter(base, sigma=3.0)
merged = 0.50 * ms1 + 0.35 * ms2 + 0.15 * ms3
merged *= brain_mask.astype(np.float32)
vals = merged[brain_mask]
if vals.size == 0:
return np.zeros_like(merged, dtype=np.float32)
support_thr = float(np.quantile(vals, 0.58))
support = merged >= support_thr
support &= brain_mask
support = ndi.binary_closing(support, structure=np.ones((5, 5, 5), dtype=bool), iterations=2)
support = ndi.binary_dilation(support, structure=np.ones((3, 3, 3), dtype=bool), iterations=1)
support = ndi.binary_fill_holes(support)
labeled, num_cc = ndi.label(support)
if num_cc > 0:
min_size = max(32, int(brain_mask.sum() * 0.0012))
keep_mask = np.zeros_like(support, dtype=bool)
for idx in range(1, num_cc + 1):
comp = labeled == idx
if int(comp.sum()) >= min_size:
keep_mask |= comp
support = keep_mask
support_soft = ndi.gaussian_filter(support.astype(np.float32), sigma=1.1)
display = merged * np.clip(support_soft, 0.0, 1.0)
display *= brain_mask.astype(np.float32)
supersample = max(1, int(supersample))
if supersample > 1:
up = ndi.zoom(display, zoom=(supersample, supersample, supersample), order=3)
up = ndi.gaussian_filter(up, sigma=max(0.1, blur_sigma * 0.8))
down_zoom = tuple(float(s) / float(u) for s, u in zip(display.shape, up.shape))
display = ndi.zoom(up, zoom=down_zoom, order=3)
display = _resize_to_match(display, attn01.shape)
display = ndi.gaussian_filter(display, sigma=max(0.1, blur_sigma))
display *= brain_mask.astype(np.float32)
vals2 = display[brain_mask]
if vals2.size > 0:
floor_q = float(np.quantile(vals2, 0.18))
display = np.clip(display - floor_q, 0.0, None)
return _normalize_01(display)
def _generate_smoothgrad_map(
model: torch.nn.Module,
image: torch.Tensor,
target_label: int,
samples: int,
noise_std: float,
) -> np.ndarray:
model.eval()
input_std = float(image.detach().std().item())
abs_sigma = max(1e-4, noise_std * max(input_std, 1e-2))
grad_acc = torch.zeros_like(image)
for _ in range(max(1, int(samples))):
noise = torch.randn_like(image) * abs_sigma
noisy = (image + noise).detach().requires_grad_(True)
logits = model(noisy)
score = logits[:, target_label].sum()
model.zero_grad(set_to_none=True)
score.backward()
if noisy.grad is None:
continue
grad_acc += (noisy.grad * noisy).detach().abs()
sal = grad_acc / float(max(1, int(samples)))
sal = sal[0, 0].detach().cpu().numpy().astype(np.float32)
return _normalize_01(sal)
def _coarse_reconstruct_for_display(attn01: np.ndarray, brain_mask: np.ndarray, downsample: int) -> np.ndarray:
downsample = max(1, int(downsample))
base = attn01 * brain_mask.astype(np.float32)
if downsample > 1:
low = ndi.zoom(base, zoom=(1.0 / downsample, 1.0 / downsample, 1.0 / downsample), order=1)
low = ndi.gaussian_filter(low, sigma=1.0)
zoom_back = tuple(float(t) / float(s) for t, s in zip(base.shape, low.shape))
rec = ndi.zoom(low, zoom=zoom_back, order=3)
rec = _resize_to_match(rec, base.shape)
else:
rec = base
rec = ndi.gaussian_filter(rec, sigma=2.0)
rec *= brain_mask.astype(np.float32)
return _normalize_01(rec)
def _resize_to_match(src: np.ndarray, target_shape: Tuple[int, int, int]) -> np.ndarray:
if src.shape == target_shape:
return src.astype(np.float32, copy=False)
zoom = [t / s for s, t in zip(src.shape, target_shape)]
return ndi.zoom(src, zoom=zoom, order=1).astype(np.float32)
def _resolve_nii_path(base_or_file: str, nii_name: str) -> str:
base_or_file = str(base_or_file).strip()
nii_name = str(nii_name).strip()
if not base_or_file:
return ""
# If an existing directory is provided, combine it with nii_name.
if os.path.isdir(base_or_file):
return os.path.join(base_or_file, nii_name)
# If not an existing path and looks like a directory style input, also combine.
if not os.path.exists(base_or_file):
if base_or_file.endswith(os.sep) or ("." not in os.path.basename(base_or_file)):
return os.path.join(base_or_file, nii_name)
# Otherwise treat it as a direct file path.
return base_or_file
def generate_heatmap(
model: torch.nn.Module,
args: argparse.Namespace,
output_dir: str,
) -> None:
os.makedirs(output_dir, exist_ok=True)
origin_nii_file = str(args.origin_nii_file).strip()
if origin_nii_file and not os.path.isfile(origin_nii_file):
raise FileNotFoundError(f"origin_nii_file not found: {origin_nii_file}")
if not os.path.isfile(args.brain_nii_file):
raise FileNotFoundError(f"brain_nii_file not found: {args.brain_nii_file}")
brain_name = os.path.basename(args.brain_nii_file)
if origin_nii_file:
origin_name = os.path.basename(origin_nii_file)
shutil.copy2(origin_nii_file, os.path.join(output_dir, origin_name))
shutil.copy2(args.brain_nii_file, os.path.join(output_dir, f"_brain_{brain_name}"))
batch = val_transform({"image": args.brain_nii_file})
image = batch["image"].unsqueeze(0).to(args.device)
model.eval()
with torch.no_grad():
logits = model(image)
logits_np = logits.detach().cpu().numpy()
pred_label = int(np.argmax(logits_np, axis=1)[0])
target_label = pred_label if int(args.nii_label) < 0 else int(args.nii_label)
brain_nii = nib.load(args.brain_nii_file)
brain_arr = np.asarray(brain_nii.get_fdata(), dtype=np.float32)
brain_mask = _build_brain_mask(brain_arr)
attn01 = _generate_smoothgrad_map(
model=model,
image=image,
target_label=target_label,
samples=args.smoothgrad_samples,
noise_std=args.smoothgrad_noise_std,
)
attn01 = _resize_to_match(attn01, brain_arr.shape)
attn01 = _coarse_reconstruct_for_display(attn01, brain_mask, args.display_downsample)
attn01_masked = attn01 * brain_mask.astype(np.float32)
clip_min = float(args.display_clip_min)
clip_max = float(args.display_clip_max)
attn01_masked = np.clip(attn01_masked, clip_min, clip_max)
attn01_masked = (attn01_masked - clip_min) / max(clip_max - clip_min, 1e-8)
attn01_masked *= brain_mask.astype(np.float32)
case_name = _strip_nii_ext(brain_name)
raw_path = os.path.join(output_dir, f"{case_name}_heatmap_raw_01.nii.gz")
nib.save(
nib.Nifti1Image(attn01_masked.astype(np.float32), brain_nii.affine, brain_nii.header),
raw_path,
)
np.save(os.path.join(output_dir, "prediction_logits.npy"), logits_np)
with open(os.path.join(output_dir, "heatmap_meta.txt"), "w", encoding="utf-8") as f:
f.write(f"model_name={args.model_name}\n")
f.write(f"model_path={args.model_path}\n")
f.write(f"origin_nii_file={args.origin_nii_file}\n")
f.write(f"brain_nii_file={args.brain_nii_file}\n")
f.write(f"heatmap_method=smoothgrad\n")
f.write(f"smoothgrad_samples={args.smoothgrad_samples}\n")
f.write(f"smoothgrad_noise_std={args.smoothgrad_noise_std}\n")
f.write(f"display_downsample={args.display_downsample}\n")
f.write(f"pred_label={pred_label}\n")
f.write(f"target_label={target_label}\n")
f.write(f"heatmap_raw_path={raw_path}\n")
f.write(f"display_blur_sigma={args.display_blur_sigma}\n")
f.write(f"display_supersample={args.display_supersample}\n")
f.write(f"display_clip_min={args.display_clip_min}\n")
f.write(f"display_clip_max={args.display_clip_max}\n")
def parse_args() -> argparse.Namespace:
parser = argparse.ArgumentParser(description="Generate ours model heatmap with medcam")
parser.add_argument("--model_name", type=str, default="SegTom", help="model_name")
parser.add_argument("--num_classes", type=int, default=3, help="number of classes")
parser.add_argument(
"--model_path",
type=str,
default="/leixin/cv/Template/run/new-segtom/best.pth",
help="path to trained model checkpoint",
)
parser.add_argument(
"--nii_name",
type=str,
default="007_S_0041_2010-01-14.nii.gz",
help="nii file name for the heatmap generation",
)
parser.add_argument(
"--origin_nii_file",
type=str,
default="",
help="original nii file path or directory; if directory, it will be joined with --nii_name",
)
parser.add_argument(
"--brain_nii_file",
type=str,
default="",
help="brain nii file path or directory; if directory, it will be joined with --nii_name",
)
parser.add_argument(
"--nii_label",
type=int,
default=-1,
help="target class label, use -1 to auto use predicted label",
)
parser.add_argument("--device", type=str, default="cpu", help="cpu or cuda")
parser.add_argument(
"--output_dir",
type=str,
default="",
help="output directory; default: outputs/hotmap_<case_name>",
)
parser.add_argument(
"--smoothgrad_samples",
type=int,
default=24,
help="number of noisy samples for SmoothGrad heatmap",
)
parser.add_argument(
"--smoothgrad_noise_std",
type=float,
default=0.14,
help="relative noise std for SmoothGrad",
)
parser.add_argument(
"--display_downsample",
type=int,
default=4,
help="downsample factor for coarse display reconstruction",
)
parser.add_argument(
"--display_blur_sigma",
type=float,
default=1.6,
help="gaussian blur sigma for display heatmap smoothing",
)
parser.add_argument(
"--display_supersample",
type=int,
default=2,
help="super-sampling factor for display interpolation smoothing",
)
parser.add_argument(
"--display_clip_min",
type=float,
default=0.1,
help="min clip value on normalized heatmap before remapping to [0, 1]",
)
parser.add_argument(
"--display_clip_max",
type=float,
default=0.9,
help="max clip value on normalized heatmap before remapping to [0, 1]",
)
args = parser.parse_args()
if not (0.0 <= args.display_clip_min < args.display_clip_max <= 1.0):
raise ValueError("display_clip_min and display_clip_max must satisfy 0 <= min < max <= 1")
return args
def main() -> None:
args = parse_args()
args.origin_nii_file = _resolve_nii_path(args.origin_nii_file, args.nii_name)
args.brain_nii_file = _resolve_nii_path(args.brain_nii_file, args.nii_name)
model = select_model(args)
checkpoint = torch.load(args.model_path, map_location=args.device)
state_dict = checkpoint.get("model_state_dict", checkpoint)
model.load_state_dict(state_dict)
model.to(args.device)
case_name = os.path.basename(args.brain_nii_file)
if case_name.endswith(".nii.gz"):
case_name = case_name[:-7]
else:
case_name = os.path.splitext(case_name)[0]
output_dir = args.output_dir or os.path.join("outputs", f"hotmap_{case_name}")
generate_heatmap(
model=model,
args=args,
output_dir=output_dir,
)
if __name__ == "__main__":
main()